Dalhousie University
Halifax
, Nova Scotia
Canada
Graduate Position
Bachelor's, Masters
Funded PhD developing tools to automatically identify new and emerging antimicrobial resistance
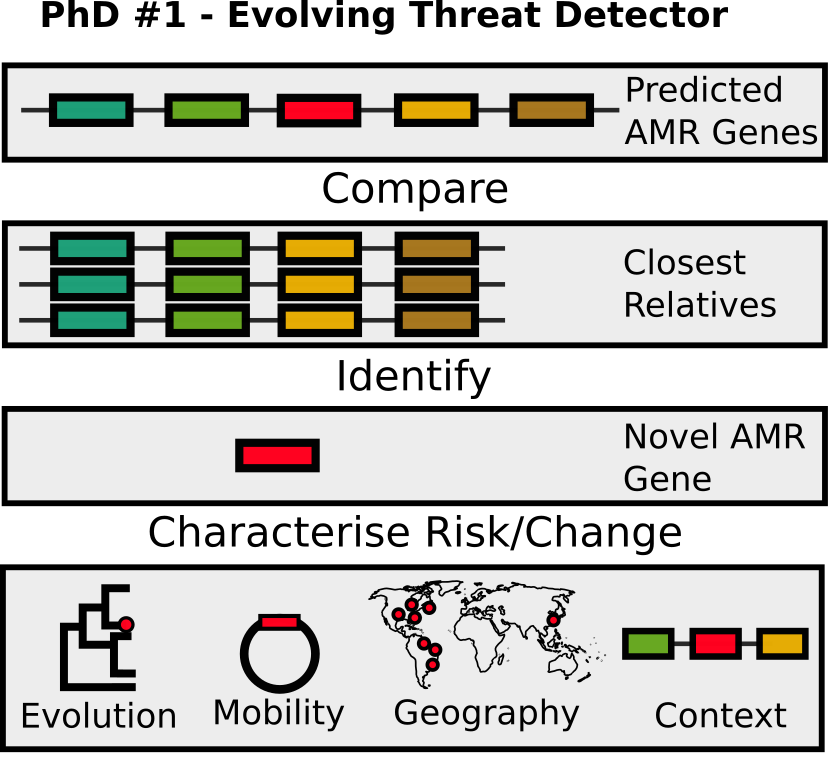
The main aim of the PhD will be the development and evaluation of methods for automatically identifying novel and evolving antimicrobial resistance (AMR) genes. AMR is a growing global health crisis that threatens the function of modern medicine. Genomically informed diagnostics and surveillance have been identified as key activities to try and ameliorate this crisis. Currently, AMR gene prediction results (such those generated by CARD’s RGI) require a lot of expertise and manual analyses in order to identify potentially important or worrying emergent genes. This project seeks to use existing data from efforts to characterise AMR in large genome databases along with associated phylogenetic, spatiotemporal, and phenotypic data to develop means of automating these analyses.
The PhD will take place within the Maguire Lab in the Faculty Computer Science & Faculty of Medicine (Department of Community Health & Epidemiology) at Dalhousie University. The specific PhD program (CS, Epidemiology, Interdisciplinary or potentially Microbiology/Biochemistry) will depend on the specific candidate background. Through this research the candidate will gain skills in the analysis of large complex datasets, the development of bioinformatic tools and web resources, and translating genomic data into clinical and public health insights.
This project will also involve existing collaborations with clinicians (Toronto’s Shared Hospital Laboratory and Sunnybrook Research Institute), public health agencies (PHAC/NML & CFIA/NCFAD), and infectious disease researchers including local experts in evolutionary microbiology and Dr. Andrew McArthur’s Comprehensive Antibiotic Resistance Database (card.mcmaster.ca) group at McMaster University.
Candidates should have:
- A Masters degree in any of Computer Science, Microbiology, Molecular Biology, Epidemiology or related fields.
- GPA of at least 3.4/4 (approx. 3.7/4.3 or or 8.6/10 or 17/20) or equivalent.
- Demonstrated fluency in English (e.g., first language, degrees taught in English, IELTS (all categories of >=7), TOEFL (>=95) or equivalent test results).
- Experience with data visualisation and analysis including familiarity with a scripting language (e.g., Python).
- Ability to work independently and within a team environment
- Effective oral and written communication, analytical, and interpersonal skills.
Prior experience in bioinformatics is not necessary but it is not desired.
We encourage all qualified applicants to apply. This lab is strongly committed to diversity within its community and especially welcomes applications from racialized persons / persons of colour, women, Indigenous / Aboriginal People of North America, persons with disabilities, LGBTQ2S+ persons, and others who may contribute to the further diversification of ideas. Our values regarding equity and diversity are linked with our commitment to excellence in the pursuit of our academic mission.
Interested individuals should send:
- Brief cover letter with most relevant experience and career goals.
- CV detailing academic training and research to date.
- Unofficial academic transcripts.
Applicants who meet preliminary screening will be invited for an interview and 2 references will be requested prior to final hiring decisions.
Please send applications to finlay.maguire@dal.ca. In the body of your email, please mention your earliest availability date.
Algorithms; Biomedical; Biostatistics; Infectious Diseases; Computational Machine Learning; Microbial Pathogenomics
Posted on: